Paper2Graph
Paper2Graph (P2G) uses GPT-derived knowledge graphs to extract, represent, and discover knowledge from scientific papers. This pilot analysis explored pharmacological use cases, comparing GPT-extracted drug pathways against gold-standard manually curated knowledge bases.
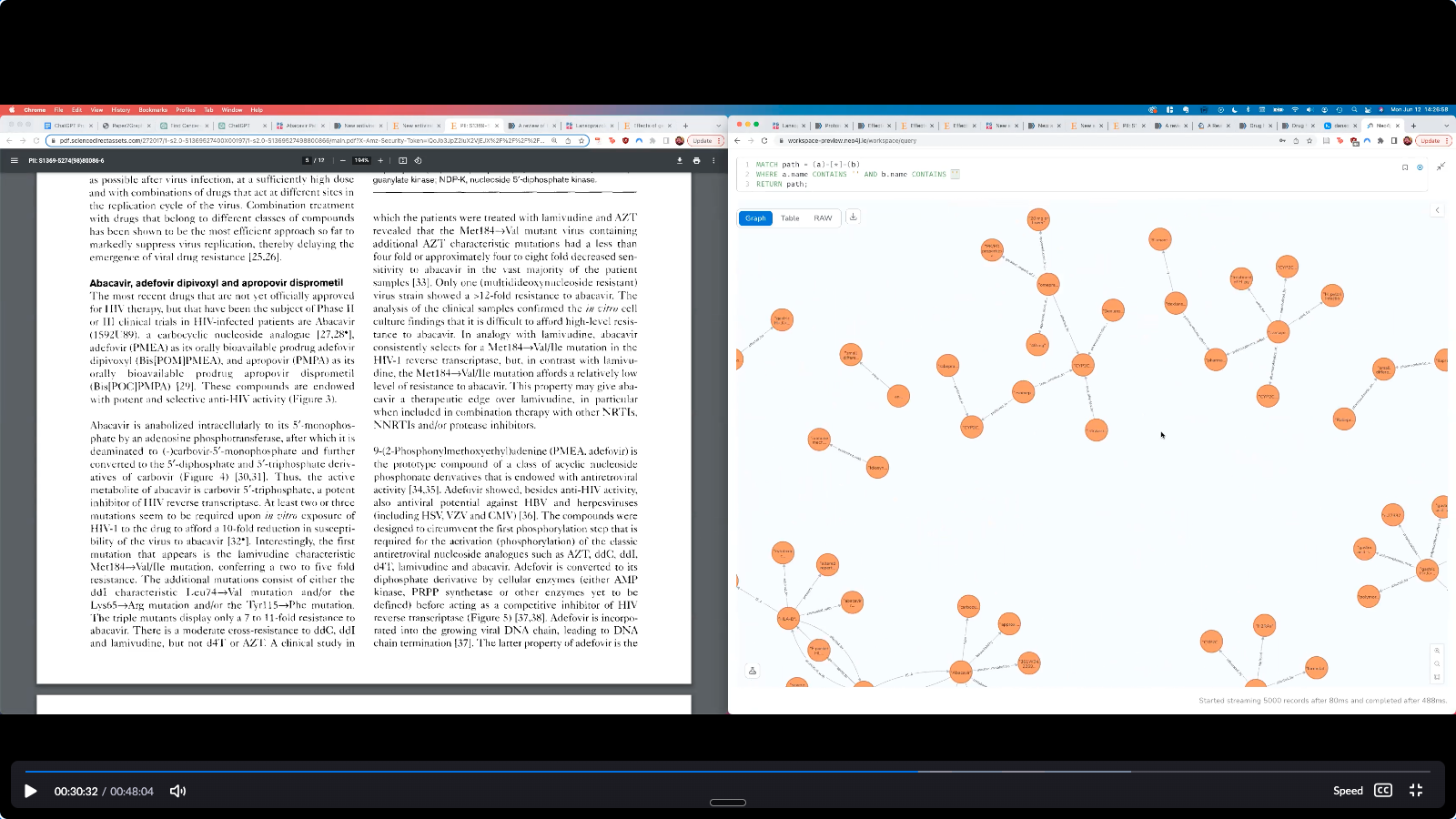
Source paper (left) and GPT-extracted knowledge graph in Neo4j (right)
Objectives
Recreate existing manually curated drug pathways and qualitatively assess knowledge extraction quality, focusing on hallucinations and mapping to biomedical ontologies.
Explore metapaths in Neo4j for recapitulating known drug mechanisms via Cypher queries.
Objective 1 – Recapitulating Gold-Standard Knowledge
Methods
PharmGKB, an authoritative manually-curated knowledge base of personalized drug response, was referenced for drug pathways. Two drug pathways were studied:
- Lansoprazole (simple pathway) – P1: pharmacokinetics & pharmacodynamics; P2: PD in children
- Abacavir (complex, multiple sub-pathways) – P1: genetic associations/adverse reactions; P2: pharmacokinetics; P3: pharmacodynamics
Text from source papers was fed into Paper2Graph. GPT-4 was used in all cases, with duplicate runs to assess non-deterministic behavior.
Observations: Strengths
- Knowledge captured with high fidelity – good sentence syntax parsing and entity identification
- Accurate extraction of quantitative data (e.g., "20mg in Japan, 40mg in US")
- Accurate extraction of biological relations: "binds to", "associated with"
- NER picks up fine-grained details like study design identifiers ("AZ0001")
Observations: Challenges
- Normalization: Equivalent entities not mapped together; singleton nodes; ambiguous acronyms (e.g., "PPIs" = Protein-Protein Interaction vs. Proton Pump Inhibitor)
- Over-broad entities: Some text spans too large to be biologically meaningful (e.g.,
CYP2C19, gastric H+,K+-ATPase genotype) - Non-determinism: Two runs on same input yield ~80% overlap – 20% variation
- Hallucination: Some fabricated triples present
- Lost context:
(Abacavir, causes, severe adverse reactions)– dangerous without the genetic variant qualifier - Inaccurate semantics:
(Drug X, associated with, genetic variant Y)– semantically close but wrong
Objective 2 – Exploring Drug Mechanism Paths
Methods
The KGs from Objective 1 were interrogated via Cypher queries to generate mechanistic understanding of drug function. Questions posed:
- By what mechanism does abacavir lead to a hypersensitivity adverse reaction?
- How does abacavir inhibit HIV?
- What is known about lansoprazole's effect on PPIs?
- What adverse reaction is this drug known to cause?
- At what dosage is this drug effective?
- Why might a new drug X be effective in this disease context? (repurposing)
- What normal function can be inhibited by a drug with no major harmful downstream consequences? (toxicology)
Observations
- Entity normalization is key for retrieving mechanistic knowledge paths – unconstrained GPT yields disconnected components
- Manual curation and knowledge updating tools are essential when using noisy GPT ingestion
- Natural language → Cypher queries is promising but challenging to interpret correctly
Conclusions
GPT-derived knowledge bases present a great opportunity to extract knowledge and rich semantics at scale. GPT-4 does well at sentence parsing, entity identification, and capturing quantitative information. Key remaining work includes ensuring high quality, reducing noise, preserving context, and tailoring output to specific end users.
Design considerations around noise tolerance, KG detail level, and pre-processing depend heavily on the end user – a toxicologist at Merck has very different needs from an academic computational pharmacologist. These tools will undoubtedly augment humans' ability to understand and contribute to science.